Quantum Control and Sensing by Spin Cooling
To reveal “invisible” NMR signal of surfaces, active sites, and functional species in catalysis, molecular recognition and quantum materials using out of the box tools.
The Han Lab pushes the frontiers of magnetic resonance and quantum information science with the goal of “seeing” chemical and biological building blocks and processes at the quantum limit. The two main themes of the Han Lab over the past 20 years have been spins and water. Electron and nuclear spins are the ultimate quantum reporters and contrast agents for biochemical processes and chemical building blocks. Advanced magnetic resonance sensing, control over the spatial organization of electron and nuclear spin clusters, and dual electron-nuclear magnetic resonance techniques have contributed to uncovering new design rules for molecular recognition, as well as the surface structuring, shaping, and ordering of biological water.
Recently, we have begun to ask the ultimate question: do quantum phenomena direct and control biological and chemical processes? The answer is yes, but high-quality experimental validations are key to qualifying the context and boundaries of these answers. Recent breakthrough developments by the Han Lab offer one-of-a-kind experimental tools that allow us to gain control over the initialization and manipulation of quantum spin states via spin cooling at high magnetic fields.
This development effort requires interdisciplinary research tools, including instrument development to combine electron and nuclear magnetic resonance with optical excitation and detection, the design of precisely tuned electron and nuclear spin qubits, spin dynamics simulations, and studies of the dynamics and thermodynamics of solvation to control biomolecular activity and assembly.
We are motivated by the power of “Seeing is Believing.” New tools for visualizing molecular interactions and materials interfaces—previously “invisible”—have fundamentally transformed our ability to discover solutions and ask new questions. The next frontier of visualization will be quantum microscopy that relies on spins as quantum reporters to deliver molecular insights that conventional microscopy cannot.

Research in the Han Lab develops novel tools to exploit spins as quantum reporters with unprecedented sensitivity and information content and as biological qubits with spin state control. Our core interest lies in advancing spin-based quantum information science, solvation science, and the molecular basis of signal transduction.
SEE ALL RESEARCHTo reveal “invisible” NMR signal of surfaces, active sites, and functional species in catalysis, molecular recognition and quantum materials using out of the box tools.
To reveal long-standing questions on the structure and dynamics of water on proteins, membranes to catalyst support surfaces.
To understand, control and engineer protein aggregation pathways, protein surface activity to protein liquid-liquid phase separation.
A. S. Roy, K. Tsay, P. P. Borbat, A. Destefano, S. Han, M. Srivastava, J. H. Freed. J. Am. Chem. Soc., 2026, 148, 2378-2387. https://doi.org/10.1021/jacs.5c16298
Corrections J. Am. Chem. Soc. 2026, 148, 8, 9140,. https://doi.org/10.1021/jacs.6c02082
Intrinsically disordered proteins (IDPs) underlie essential cellular functions and drive neurodegenerative diseases through mutation-induced structural changes, yet their conformational heterogeneity often evades crystallography and cryo-EM. Electron spin resonance (ESR) pulsed dipolar spectroscopy (PDS), which determines distance distributions between a pair of spin-labeled residues in a protein, can provide complementary and meaningful information related to conformational heterogeneity in IDPs. Double quantum coherence (DQC) is an important ESRPDS technique, capable of measuring a wide range of distances(∼10toat least80Å), and is a single-frequency technique with a small back ground that can be easily removed. This makes DQC an ideal candidate to probe IDPs. We present a complete theoretical framework for DQC data analysis, incorporating pseudo-secular dipolar coupling and finite pulse effects, enabling rapid and accurate reconstruction of complex distance distributions in doubly nitroxide-labeled IDPs. We validate the method on rigid biradicals with known inter-spin distances. The application to a tau protein fragment(jR2R3) reveals distinct end-to-end distance distributions for the wild-type vs the disease-associatedP301Lmutant. The results expose differences in their conformationaldistributions,whichlikelygoverntheirdivergentaggregationpropensities.ThisadvancealsoestablishesDQCESR as a powerful, accessible tool for probing disorders in biomolecular systems.
Q. Stern, J. Cui, R. Chaklashiya, C. Tobar, M. Judd, O. Nir-Arad, D. Shimon, I. Kaminker, H. Takahashi, J. R. Sirigiri, and S. Han Phys. Chem. Chem. Phys., 2026. https://doi.org/10.1039/D5CP04594K
We investigate nitrogen substitution defects, also known as P1 centers, in Type 1b diamonds generated under high pressure and high temperature (HPHT) as a source of electron spin polarization. The open question was how readily electron spin polarization in this diamond transfers to 13C nuclear spins within the diamond matrix at 14 T by dynamic nuclear polarization (DNP). The goal was to refine the model for clustered P1 centers in HPHT diamonds and evaluate their potential as a source for DNP hyperpolarization or contrast. The study relied on frequency-stepped measurements of DNP profiles under magic angle spinning (MAS) using the mm-wave output of a frequency-tunable gyrotron and a regular superconducting NMR magnet at a single field. We observe up to 700-fold 13C on/off signal enhancements in both MAS and static mode at room temperature, and 130-fold between 35 and 100 K. Modelling the experimental results revealed the dominant role of P1 clusters harboring inter-P1 dipolar and exchange couplings exceeding 100 MHz in achieving 13C DNP at 14.1 T. Our results exemplify the importance of exchange coupling for high-field DNP and provides a refined model for DNP via P1 centers of HPHT diamonds.
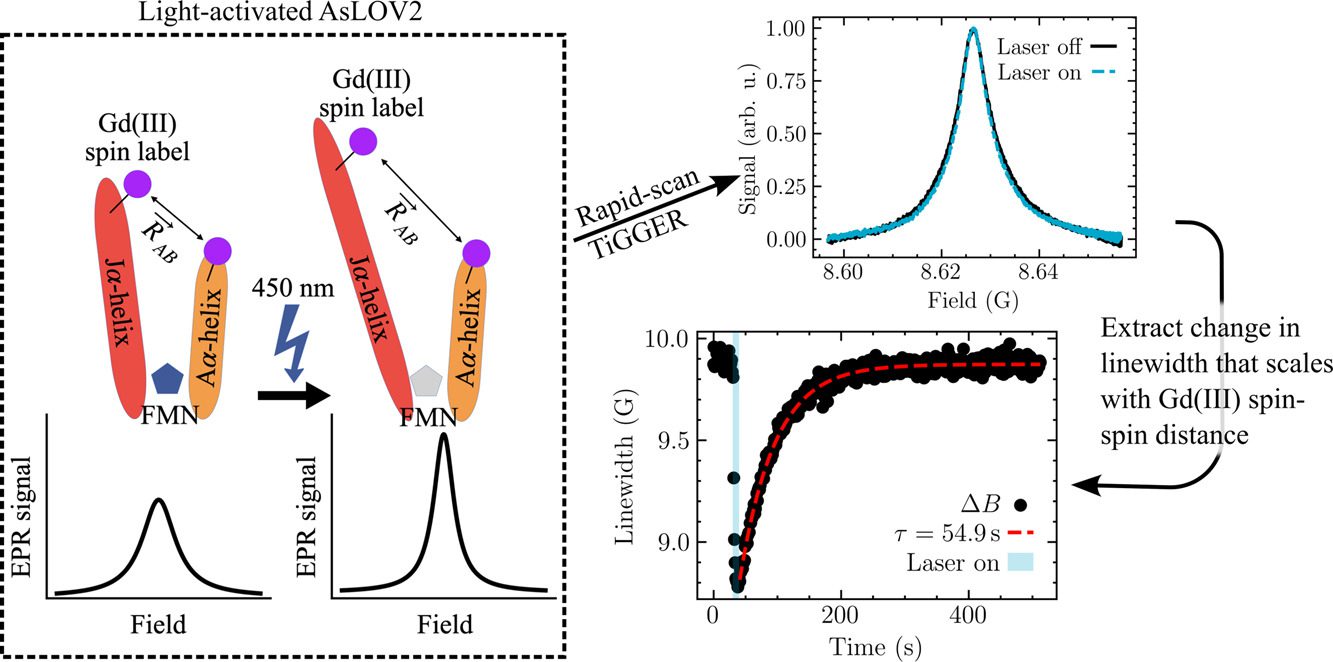
We present field-domain rapid-scan (RS) electron paramagnetic resonance (EPR) at 8.6 T and 240 GHz. To enable this technique, we upgraded a homebuilt EPR spectrometer with an FPGA-enabled digitizer and real-time processing software. The software leverages the Hilbert transform to recover the in-phase (𝐼) and quadrature (𝑄) channels, and therefore the raw absorptive and dispersive signals, 𝜒′ and 𝜒′′, from their combined magnitude (√𝐼2+𝑄2). Averaging a magnitude is simpler than real-time coherent averaging and has the added benefit of permitting long-timescale signal averaging (up to at least 2.5 × 106 scans) because it eliminates the effects of sourcereceiver phase drift. Our rapid-scan (RS) EPR provides a signal-to-noise ratio that is approximately twice that of continuous wave (CW) EPR under the same experimental conditions, after scaling by the square root of acquisition time. We apply our RS EPR as an extension of the recently reported time-resolved Gd-Gd EPR (TiGGER) [Maity et al., 2023], which is able to monitor inter-residue distance changes during the photocycle of a photoresponsive protein through changes in the Gd-Gd dipolar couplings. RS, opposed to CW, returns field-swept spectra as a function of time with 10 ms time resolution, and thus, adds a second dimension to the static field transients recorded by TiGGER. We were able to use RS TiGGER to track time-dependent and temperature-dependent kinetics of AsLOV2, a light-activated phototropin domain found in oats. The results presented here combine the benefits of RS EPR with the improved spectral resolution and sensitivity of Gd chelates at high magnetic fields. In the future, field-domain RS EPR at high magnetic fields may enable studies of other real-time kinetic processes with time resolutions that are otherwise difficult to access in the solution state.